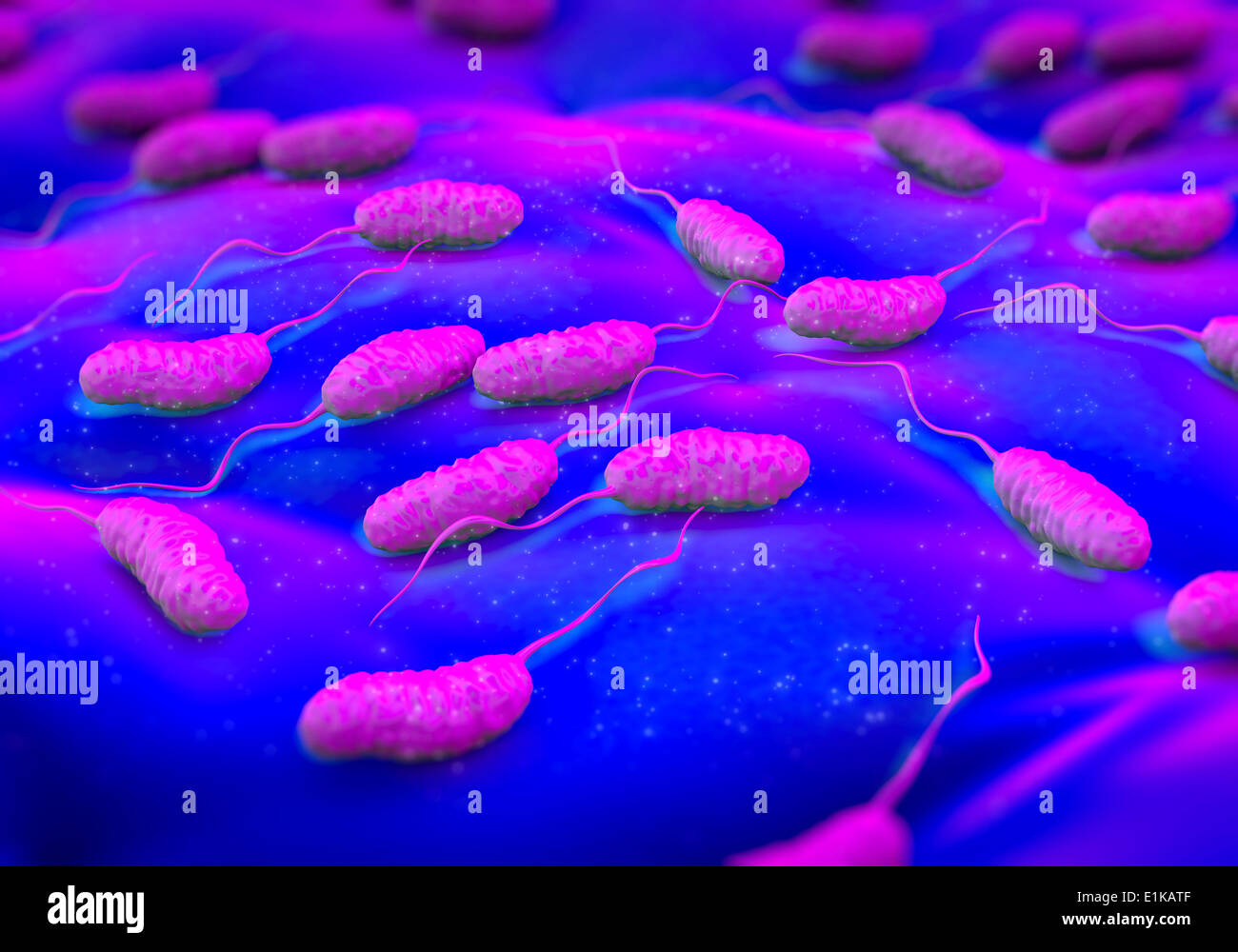
GGA AAG ATC TAT GCG TCG ACG TTA CCG ATG CTA TGG GĬAT ATG GAA TTC CCG GGA TCC ATG CTC TAG AAG TCG GTT GTT TCG GTA A

The high antibiotic resistance and virulence gene polymorphism suggested that it is important to strengthen the surveillance for the genetic variation and drug resistance of the clinical isolates of non-O1/O139 V.cholerae. The predominant virulence pattern was hlyA- rtxA- hapA- ompU- nanH- vasA- vasK- vasH- vcsC- vcsV- vcsN- vspD (40.0%). PCR detection of virulence genes showed that all strains were positive for hlyA and ompU gene, and the positive rates of hapA, rtxA, T6SS, T3SS and nanH were 97.1%, 91.4%, 94.3%-97.1%, 80.0%-85.7%, 62.9%, respectively. The strains were sensitive to amoxicillin/clavulanate, ceftriaxone, cefoxitin, cefepime and imipenem. The drug susceptibility test showed that the resistant rate of 35 strains of non-O1/O139 V.cholerae to trimethoprim-sulfamethoxazole was highest (40.0%), followed by the resistant rates to chloramphenicol (28.5%) and sulfisoxazole (22.6%).
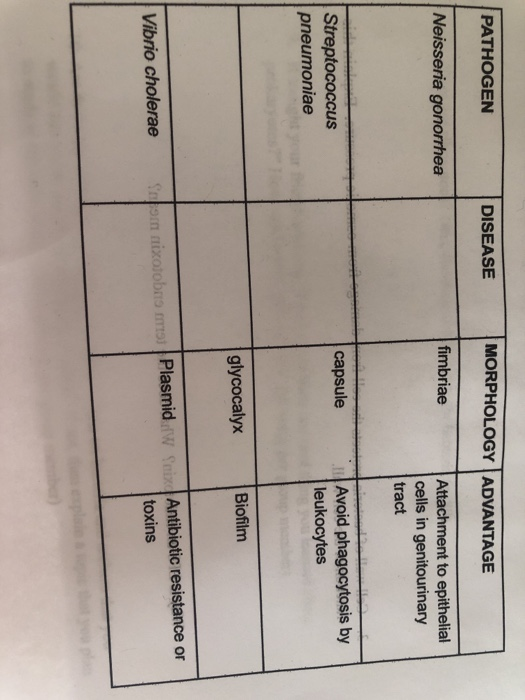
The susceptibility of V.cholerae strain to 15 antibiotics was tested with microbroth dilution method, and virulence genes were detected by using polymerase chain reaction (PCR). Non-O1/O139 V.cholerae strains and the related patient records were collected from the intestinal clinic of Beijing Friendship Hospital from 2014 to 2015. To explore the distribution of virulence genes and drug resistance of non-O1/O139 Vibrio (V.) cholerae.
